| | ||||||||||
| 1 | INN | Class | Route (list) | PK parameters= Cmax; Tmax; F: bioavailability; t1/2: half-life; VD: volume of distribution; Cl: clearance; PPB: plasma protein binding;(EQN means that the equation t1/2 = VD / Cl * 0.693 was used | Primary Target and PDB code of Protein-Drug complex | Targets from DrugCentral | Links | |||
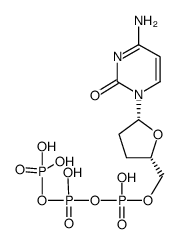
| M ZALCITABINE (is an active metabolite)
| ZALCITABINE | ANTIINFECTIVES | - | HIV REVERSE TRANSCRIPTASE (GAG-POL) DNA POLYMERASE-BETA MITOCHONDRIAL DNA POLYMERASE-GAMMA PDB 4DQP (TERNARY COMPLEX OF BACILLUS DNA POLYMERASE I LARGE FRAGMENT, DNA DUPLEX, AND DDCTP (PAIRED WITH DG OF TEMPLATE)) LIGAND CODE = DCT (link to the list of PDB complexes) Download experimental 3D coordinates of DCT with added hydrogens | Reverse transcriptase,RNaseH UNIPROT Q72547 pol more at DrugCentral | EMA | ||||
| 2 | INN | Class | Route (list) | PK parameters= Cmax; Tmax; F: bioavailability; t1/2: half-life; VD: volume of distribution; Cl: clearance; PPB: plasma protein binding;(EQN means that the equation t1/2 = VD / Cl * 0.693 was used | Primary Target and PDB code of Protein-Drug complex | Targets from DrugCentral | Links | |||
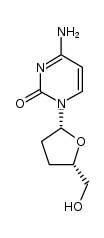
| ZALCITABINE DISCONTINUED (has an active metabolite)
| ZALCITABINE | ANTIINFECTIVES | ORAL | Cmax 119.4 NANOMOLAR Tmax 0.8 HOUR F 80 PERCENT VD 34.7 LITER (65 KILOGRAM) Cl 17.1 LITER / HOUR HT 2 HOUR SOLUBILITY 76.4 MILLIGRAM / MILLILITER IN WATER | DEOXYCYTIDINE KINASE PDB 2VP9 (STRUCTURAL STUDIES OF NUCLEOSIDE ANALOG AND FEEDBACK INHIBITOR BINDING TO DROSOPHILA MELANOGASTER MULTISUBSTRATE DEOXYRIBONUCLEOSIDE KINASE) LIGAND CODE = DOC (link to the list of PDB complexes) Download experimental 3D coordinates of DOC with added hydrogens | Reverse transcriptase,RNaseH UNIPROT Q72547 pol more at DrugCentral | EMA | |||