| | ||||||||||
| 1 | INN | Class | Route (list) | PK parameters= Cmax; Tmax; F: bioavailability; t1/2: half-life; VD: volume of distribution; Cl: clearance; PPB: plasma protein binding;(EQN means that the equation t1/2 = VD / Cl * 0.693 was used | Primary Target and PDB code of Protein-Drug complex | Targets from DrugCentral | Links | |||
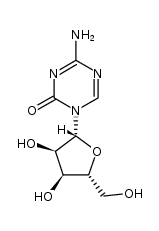
| AZACITIDINE (has an active metabolite)
| AZACITIDINE | ANTINEOPLASTIC | INTRAVENOUS SUBCUTANEOUS ORAL | Cmax 3 MICROMOLAR F 89 PERCENT VD 76 LITER Cl 167 LITER / HOUR HT 0.68 HOUR SOLUBILITY SPARINGLY SOLUBLE IN WATER | URIDINE-CYTIDINE KINASE 2 (UCK2) PDB 4QD3 (CRYSTAL STRUCTURE OF PEPTIDYL-TRNA HYDROLASE FROM PSEUDOMONAS AERUGINOSA WITH 5-AZACYTIDINE AT 1.89 ANGSTROM RESOLUTION) LIGAND CODE = 5AE (link to the list of PDB complexes) Download experimental 3D coordinates of 5AE with added hydrogens | DNA (cytosine-5)-methyltransferase 1 UNIPROT P26358 DNMT1 -- DNA (cytosine-5)-methyltransferase 3A UNIPROT Q9Y6K1 DNMT3A more at DrugCentral | EMA | |||